It began not with a sweeping hypothesis, but with a focused question. During her MSc dissertation, Miss Sakhaviya was examining how commonly used antibiotics influence the structure and ligand-binding function of human serum albumin, the principal protein responsible for transporting drugs in the bloodstream. The work was grounded in a precise objective: to understand how structural perturbations in this protein might alter its ability to bind and carry molecules. As the results started to take shape, our discussions repeatedly returned to a simple clinical reality. These antibiotics are rarely prescribed in isolation. In routine practice, they are often administered alongside medications for acidity, particularly proton pump inhibitors such as pantoprazole. That observation gradually shifted the direction of the study. Instead of focusing on a single drug–protein interaction, the question evolved into something more representative of real-world conditions: what happens when multiple drugs interact simultaneously with the same protein?
“Drugs rarely act alone within the body, they meet on shared proteins, where subtle interactions can reshape their behavior and impact.”
To approach this, it became necessary to consider the structural sensitivity of human serum albumin itself. Its function depends on a finely maintained three-dimensional conformation, and even subtle disturbances can influence how it binds drugs, how long those drugs remain in circulation, and how effectively they reach their targets. If one drug alters this structure, the implications extend beyond its own transport, potentially affecting the behavior of others present in the system. Within this context, we began to ask whether pantoprazole, beyond its primary role in reducing gastric acidity, could influence the protein against such disturbances.
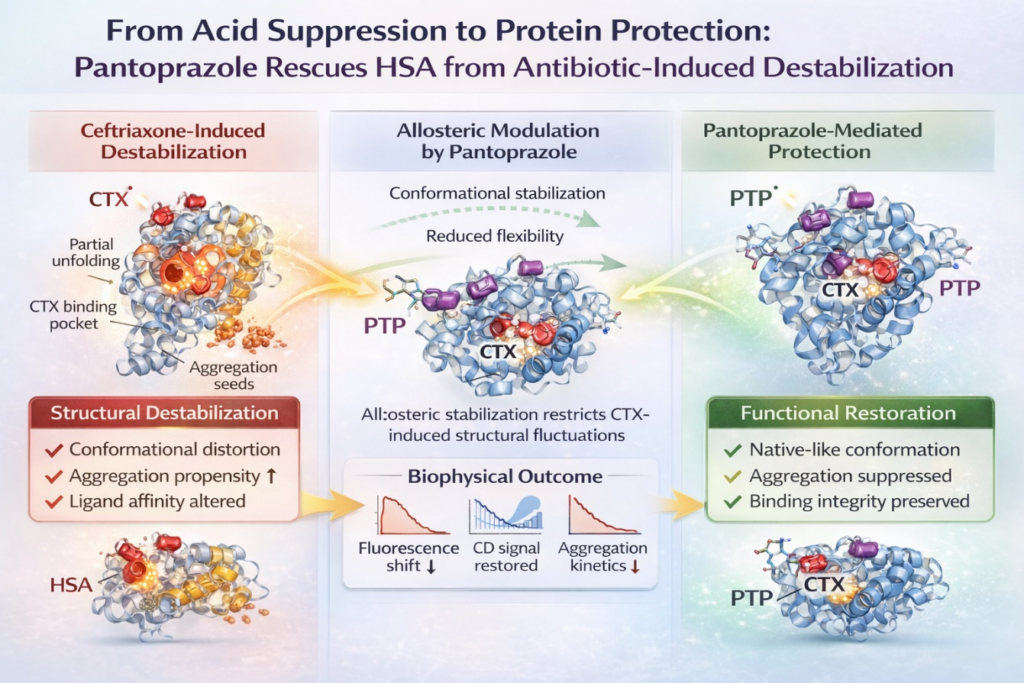
What emerged from the data was not an obvious interaction, but a consistent one. Ceftriaxone induced structural perturbations associated with unstability and aggregation. In contrast, pantoprazole appeared to counterbalance these effects, restoring features of the protein’s native conformation. The two drugs did not compete for the same binding site, suggesting that the effect was not driven by direct displacement. Instead, the observations pointed toward an allosteric mode of stabilization, where binding at one region of the protein subtly influences its overall structural landscape. The strength of this finding lay in its reproducibility across different approaches. Spectroscopic analyses indicated preservation of structural elements characteristic of native albumin. Aggregation studies showed a clear reduction in self-association under conditions where destabilization would otherwise be expected. Computational modeling supported these observations, suggesting interactions consistent with a stabilizing influence at the molecular level. Each method offered a different perspective, yet all converged on the same conclusion.
What was particularly striking was not only the presence of this effect, but its consistency. Pantoprazole is generally regarded as a supportive medication, prescribed to manage acidity rather than to influence protein behavior. Yet in this setting, it appeared to play a more active role, contributing to the maintenance of protein stability in the presence of a destabilizing agent. The observation does not redefine its primary function, but it does expand how its role can be understood at the molecular level. This brings attention to a broader point. Drug combinations are typically evaluated in terms of clinical outcomes, but their molecular interplay often remains insufficiently explored. Proteins such as human serum albumin act as shared platforms where multiple drugs converge. Interactions occurring at this level need not be competitive to be meaningful. Changes in protein conformation can indirectly influence how different molecules bind, dissociate, or redistribute within the system. Such effects are unlikely to be immediately visible in standard clinical settings. They do not necessarily produce overt signals, yet they can shape the biochemical environment in subtle ways. The findings here suggest that co-administered drugs may influence each other indirectly through shared biological targets, modifying protein structure and, in turn, affecting drug behavior. The work does not attempt to generalize beyond its scope, but it does point toward an additional layer of interaction that deserves closer attention. Understanding how drugs collectively influence protein stability could provide deeper insight into variability in therapeutic response and open new directions for studying combination treatments.