When most people hear the term diabetes, they immediately associate it with “high blood sugar.” While this is not incorrect, it is only the surface of a much deeper and more complex condition. At its core, diabetes arises from problems related to insulin, a hormone produced by the pancreas that regulates how the body uses glucose for energy. However, the real story extends far beyond insulin deficiency or resistance. Over time, elevated glucose levels trigger a cascade of biochemical changes: Blood vessels become damaged, Kidneys suffer progressive injury, Eyes develop retinal damage, Nerves deteriorate. Pancreatic β-cells are responsible for insulin secretion, but they are surprisingly fragile. Unlike many other cells, β-cells have limited capacity to regenerate. Prolonged stress caused by high glucose, lipids, and inflammation leads to their dysfunction and eventual death or by the time up to 50% of β-cell function may already be lost. Modern medicine offers various treatments, insulin therapy, oral drugs, and lifestyle interventions. However, targeting only blood glucose may not be enough. The goal is shifting from “controlling sugar” to restoring systemic balance.
Balanced diet, Regular physical activity, Weight management, these approaches improve insulin sensitivity and delay disease progression. Thus, while essential, lifestyle modification alone is rarely sufficient in established T2DM. Current therapeutic strategies for Type 2 Diabetes Mellitus (T2DM) primarily focus on controlling blood glucose levels through a combination of lifestyle modifications and pharmacological interventions. Among oral drugs, Metformin remains the first-line therapy due to its ability to reduce hepatic glucose production and improve insulin sensitivity; however, it is associated with gastrointestinal side effects, is contraindicated in severe renal impairment, and does not prevent progressive β-cell dysfunction.
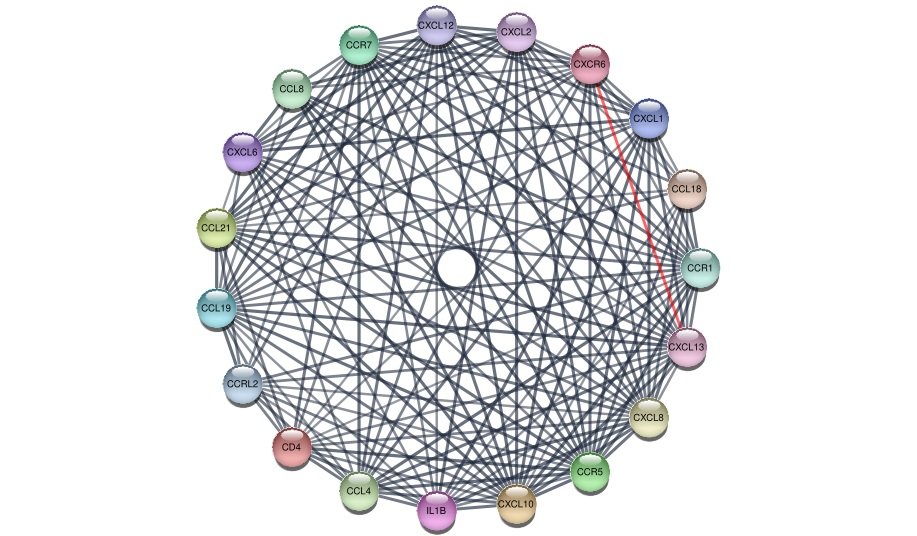
OMICS Technologies: Unraveling the Root Causes of Type 2 Diabetes The transition from symptom-based management to cause-driven therapy in Type 2 Diabetes Mellitus (T2DM) is largely powered by advances in OMICS technologies, which provide a comprehensive understanding of disease biology at multiple molecular levels. Genomics focuses on the study of genetic variations that predispose individuals to diabetes, revealing that certain genes influence insulin secretion, β-cell survival, and metabolic regulation. Transcriptomics examines gene expression patterns, offering insights into which genes are actively involved during disease progression; recent findings highlight that pancreatic β-cells may lose their functional identity under stress, rather than simply undergoing cell death. Proteomics explores the protein landscape of cells and tissues, uncovering functional disruptions such as increased inflammatory mediators and altered signaling proteins, which directly contribute to insulin resistance and cellular dysfunction. Metabolomics analyzes small-molecule metabolites, providing a real-time snapshot of metabolic changes; specific metabolites like altered lipids and amino acids have been identified as early indicators of diabetes, often detectable years before clinical symptoms arise. Beyond these individual layers, the integration of all these datasets through multi-omics approaches enables a systems-level understanding of T2DM, allowing researchers to map complex biological networks, identify key regulatory hubs, and discover novel therapeutic targets. Collectively, these methodologies are reshaping diabetes research by shifting the focus from managing elevated glucose levels to understanding and targeting the fundamental molecular mechanisms driving the disease.
OMICS and Precision Medicine: Toward Personalized Diabetes Care
The integration of OMICS technologies into diabetes research is laying the foundation for precision medicine, where treatment is tailored to an individual’s unique molecular profile rather than following a generalized approach. By combining insights from genomics, transcriptomics, proteomics, and metabolomics, researchers can now classify patients into distinct biological subtypes based on their underlying disease mechanisms such as predominant insulin resistance, β-cell dysfunction, or inflammation-driven pathology. This stratification enables clinicians to select therapies that are more likely to be effective for a specific patient, improving outcomes while minimizing adverse effects. Additionally, OMICS-driven approaches are facilitating the identification of novel, highly specific drug targets, allowing for the development of therapies that act precisely at the site of dysfunction, such as pancreatic islets or inflammatory pathways. Another key advancement is the potential for early diagnosis and intervention, as molecular alterations can be detected long before clinical symptoms appear, enabling preventive strategies tailored to individual risk profiles. Ultimately, OMICS is transforming diabetes care from a reactive, one-size-fits-all model into a proactive, personalized, and mechanism-based approach, bringing us closer to more effective and potentially disease-modifying treatments.
From OMICS to Drug Discovery: Precision Targeting through SBDD and LBDD
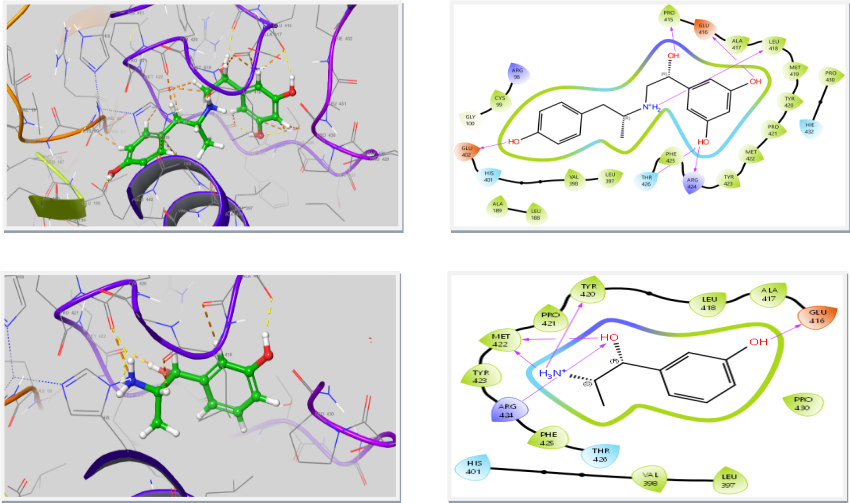
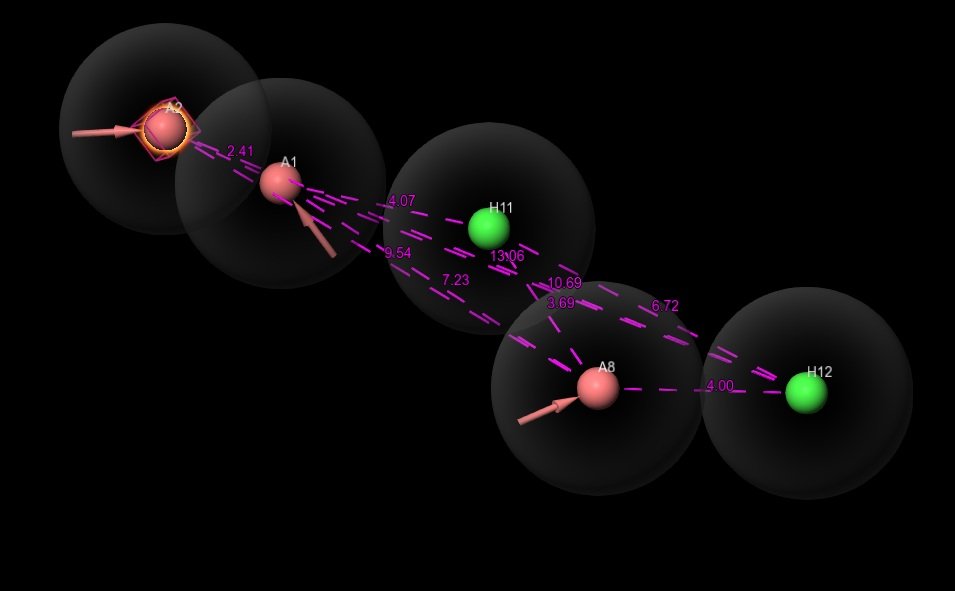
Once OMICS analyses identify key hub genes, proteins, or pathways driving Type 2 Diabetes Mellitus (T2DM), the next critical step is translating these findings into targeted therapeutic strategies through advanced drug design approaches. Two major methodologies dominate this phase: Structure-Based Drug Design and Ligand-Based Drug Design. SBDD relies on the three-dimensional structure of target proteins, enabling techniques such as molecular docking, molecular dynamics simulations, and structure optimization to predict how a drug molecule can precisely bind and modulate a specific target, thereby increasing accuracy and reducing off-target effects. In contrast, LBDD is applied when structural information is limited and focuses on known active compounds to design new molecules using approaches like QSAR modeling, pharmacophore mapping, and similarity-based screening. Together, these strategies enhance the efficiency of drug discovery by narrowing down potential candidates to those most likely to interact effectively with disease-specific targets identified through OMICS. This integration creates a highly refined pipeline moving from large-scale molecular data to precise therapeutic design ultimately improving the success rate, specificity, and translational potential of novel anti-diabetic drugs.
Why Modern OMICS-Driven Approaches Outperform Traditional Strategies
The integration of OMICS technologies with advanced drug design approaches such as Structure-Based Drug Design (SBDD) and Ligand-Based Drug Design (LBDD) offers a significant advantage over traditional trial-and-error methods of diabetes treatment and drug discovery. Conventional approaches primarily rely on generalized clinical observations and population-level responses, often focusing on symptom management, particularly glucose reduction without addressing the underlying molecular heterogeneity of Type 2 Diabetes Mellitus (T2DM). When combined with computational drug design techniques like molecular docking and QSAR modeling, significantly improves the accuracy, efficiency, and success rate of therapeutic development. Moreover, these approaches reduce time, cost, and failure rates in drug discovery by prioritizing highly probable candidates before experimental validation. Importantly, they support the development of personalized therapies, ensuring that treatments are tailored to individual patient profiles rather than relying on a one-size-fits-all model. Overall, this modern, integrated framework shifts the paradigm from reactive and generalized care to predictive, precise, and mechanism-based intervention, making it far more effective than traditional strategies.
Diabetes Research at IDDRL: Integrating OMICS and Computational Drug Design
At the IDDRL laboratory, current research efforts are focused on bridging large-scale molecular insights with precision drug discovery to address the complexities of Type 2 Diabetes Mellitus. The lab actively employs advanced OMICS approaches, including bulk RNA sequencing and single-cell RNA sequencing, to uncover differential gene expression patterns, identify disease-associated pathways, and pinpoint key regulatory hub genes involved in β-cell dysfunction, insulin resistance, and inflammatory signaling. These data-driven insights are further translated into therapeutic exploration using both Structure-Based Drug Design and Ligand-Based Drug Design methodologies. Computational platforms such as Schrödinger Suite, SYBYL-X, and GNINA are utilized for molecular docking, virtual screening, and predictive modeling to identify and optimize potential drug candidates. In parallel, the laboratory is also engaged in the discovery of novel therapeutic targets, with a particular focus on the GPR119 receptor, which plays a significant role in glucose homeostasis and insulin secretion. This integrated workflow from OMICS-driven target identification to computational drug design positions the lab at the forefront of precision-based diabetes research, aiming to develop more effective and mechanism-specific therapeutic strategies.