What first led you to study bacteria in silkworms, and how did this discovery begin?
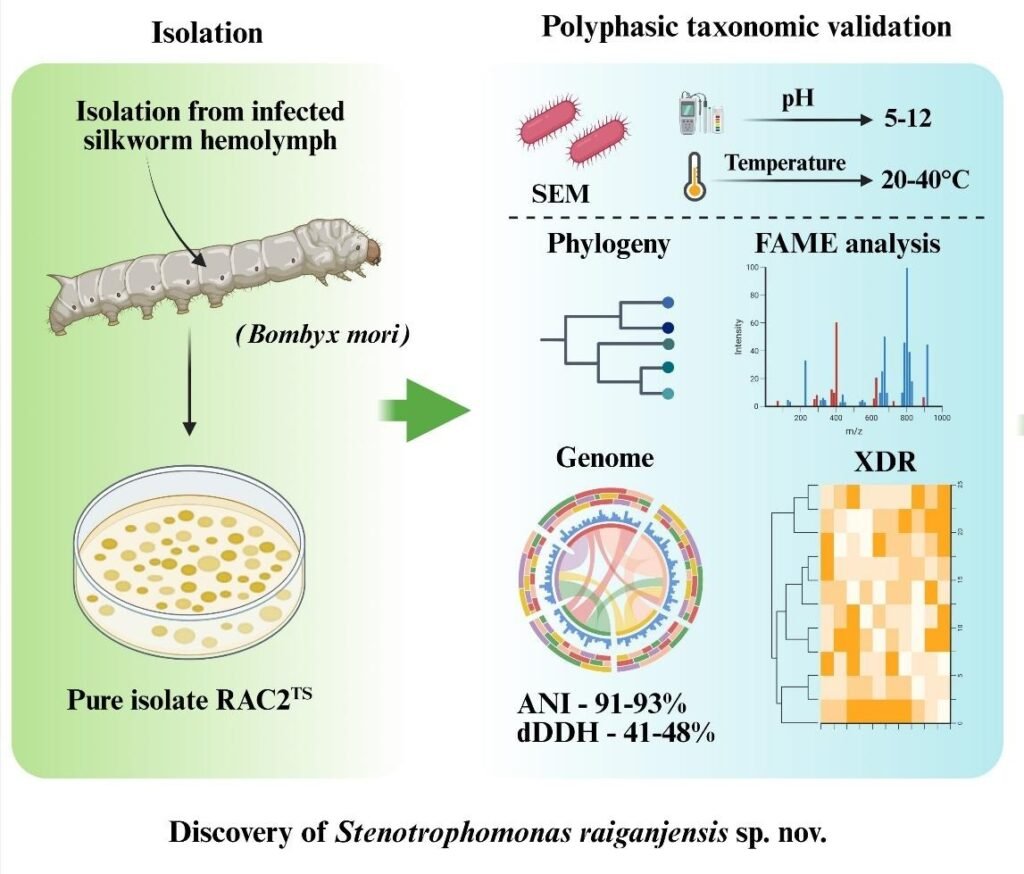
India is one of the world’s leading producers of silk, and sericulture is a crucial agro-based industry supporting millions of rural livelihoods, particularly in states such as West Bengal, Karnataka, Assam, and Tamil Nadu. Our interest in studying bacteria in silkworms originated from the frequent occurrence of bacterial diseases in rearing systems, which significantly affect sericulture productivity in India. Silkworms are highly sensitive to microbial infections, often acquired through contaminated mulberry leaves or rearing environments conditions commonly observed in traditional rearing practices. During routine screening of diseased larvae, we isolated an unusual bacterial strain from the silkworm gut. What initially appeared to be a typical infection gradually revealed distinct biological and genetic characteristics, prompting further investigation. In the Indian context, such findings are particularly important, as understanding novel bacterial associations can contribute to improved disease management strategies, enhanced rearing practices, and ultimately better silk yield and quality, thereby strengthening the sustainability of the sericulture sector.
Can you explain in simple terms what makes this bacterium new and different from others?
This bacterium is considered “new” because it shows clear differences from known species at both genetic and physiological levels. Its genome, biochemical behaviour, and growth characteristics do not fully match any previously described bacteria. In simple terms, it belongs to a known group but behaves differently enough like having a unique genetic “fingerprint” that it qualifies as a new species.
Your study mentions that this bacterium is drug-resistant, why is that important for science and society?
Drug resistance means that the bacterium can survive exposure to antibiotics that would normally kill it, and this phenomenon is fundamentally a question of molecular interactions and biochemical adaptation. Antibiotics function by targeting specific cellular components such as cell wall synthesis, protein translation, or DNA replication but drug-resistant bacteria evolve chemical strategies to evade or neutralize these effects. These may include enzymatic degradation of the drug, structural modification of antibiotic targets, altered membrane permeability, or the active efflux of antibiotics through transport proteins.
I view drug resistance as a dynamic interplay of small molecules, enzymes, and metabolic networks within the microbial system. Studying such bacteria allows us to dissect the exact chemical mechanisms underlying resistance for example, identifying resistance-conferring enzymes, characterizing metabolic shifts, or mapping interaction pathways between antibiotics and cellular targets. This insight is critical for rational drug design, enabling the development of next-generation antibiotics that can bypass resistance mechanisms or inhibit them directly.
Beyond medicine, this has broader implications for agriculture and sericulture, where resistant microbes can compromise animal health and productivity. By understanding resistance at the molecular level, we can design targeted antimicrobial compounds, develop biosensors for early detection, and create sustainable intervention strategies that minimize the emergence and spread of resistance. Thus, investigating drug-resistant bacteria is not only about combating infections but also about decoding and manipulating the chemical language of life to address one of the most pressing challenges facing science and society today.
What surprised you the most during this research or discovery process?
One of the most surprising findings was how this bacterium could persist and adapt within the silkworm gut environment while simultaneously carrying multiple drug-resistance traits. This was particularly striking because the silkworm gut, though seemingly simple, represents a highly dynamic chemical microenvironment shaped by host metabolism, dietary inputs (mulberry leaves), and microbial interactions. The bacterium’s ability to survive here suggests that it is not merely passively existing, but actively modulating its biochemical pathways adjusting metabolite production, regulating stress-response molecules, and possibly altering membrane chemistry to withstand both host defenses and antimicrobial pressures.
What stood out most was the sophistication of its survival strategies at the molecular level. The coexistence of multiple resistance traits implies coordinated chemical mechanisms, such as enzyme-mediated drug modification, efflux pump activation, and metabolic flexibility that allows the bacterium to thrive under fluctuating conditions. This level of adaptability in a niche insect gut environment challenges the conventional assumption that complex resistance evolution is restricted to clinical or heavily antibiotic-exposed settings.
This discovery highlights that even micro-scale ecosystems like the silkworm gut can function as active reservoirs and evolution hubs for chemically driven resistance mechanisms. It underscores the importance of exploring such underappreciated environments to uncover hidden biochemical diversity and resistance pathways. These insights not only deepen our understanding of microbial chemical ecology but also have significant implications for predicting resistance emergence, guiding antimicrobial design, and developing more informed strategies for both biomedical and agricultural disease management.
Do you think this bacterium could have any impact on human health, agriculture, or the environment?
Yes, potentially. Such bacteria can have significant impacts across agriculture, human health, and the environment. In agriculture, particularly in sericulture, the presence of drug-resistant bacteria in silkworms can lead to persistent infections that are difficult to control, thereby reducing cocoon yield and affecting farmer livelihoods.
From a broader chemical and ecological standpoint, these bacteria are important because they can act as reservoirs of resistance genes embedded within mobile genetic elements and biochemical pathways. Such resistance traits can spread across microbial communities through processes like horizontal gene transfer, potentially reaching human-associated pathogens which is an interconnected system driven by molecular interactions and chemical adaptation across ecosystems. Understanding how these resistance mechanisms originate, evolve, and transfer is essential not only for improving disease management in sericulture but also for predicting and mitigating the spread of antimicrobial resistance, making these bacteria highly relevant from both ecological and global public health perspectives.
What are the next steps that still need to be studied before we fully understand this bacterium?
Several important questions remain before we can fully understand this bacterium and its broader implications. First, it is essential to decipher the precise molecular and biochemical mechanisms underlying its drug resistance, including the specific genes, enzymes, and metabolic pathways involved. Second, its direct role in silkworm disease progression needs to be clearly established whether it acts as a primary pathogen, an opportunistic invader, or part of a more complex disease network. Third, understanding its interactions within the host microbiome is critical, as these microbial relationships may influence both its survival and pathogenic potential. Finally, it is important to assess any potential cross-species transmission or environmental risk, particularly in the context of resistance gene dissemination. Future research should integrate functional genomics, metabolomic profiling, controlled infection models, and ecological tracking to map how this bacterium operates at the molecular level and within real-world systems. Such studies will be crucial for building a comprehensive understanding of its biology, evolution, and impact across agricultural and environmental contexts.